 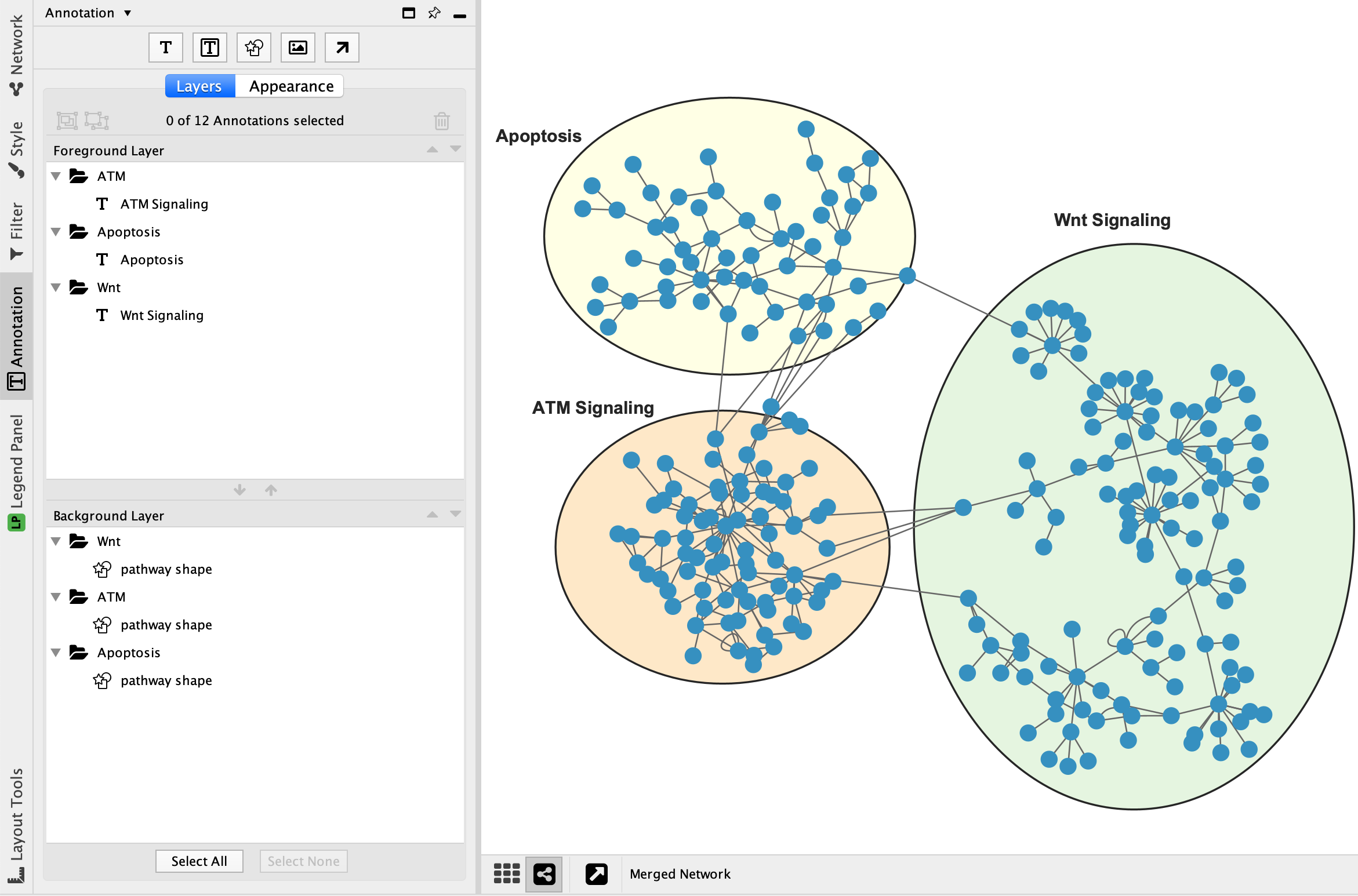
ReNE enhances a network by only integrating data from public repositories, without any inference or prediction. ReNE can be freely installed from the Cytoscape App Store ( ) and the full source code is freely available for download through a SVN repository accessible at. The enhanced network produced by ReNE is exportable in multiple formats for further analysis via third party applications. The resulting enhanced network is still a fully functional Cytoscape network where each regulatory element (transcription factor, miRNA, gene, protein) and regulatory mechanism (up-regulation/down-regulation) is clearly visually identifiable, thus enabling a better visual understanding of its role and the effect in the network behavior. The normalized network is then analyzed to include missing transcription factors, miRNAs, and proteins. 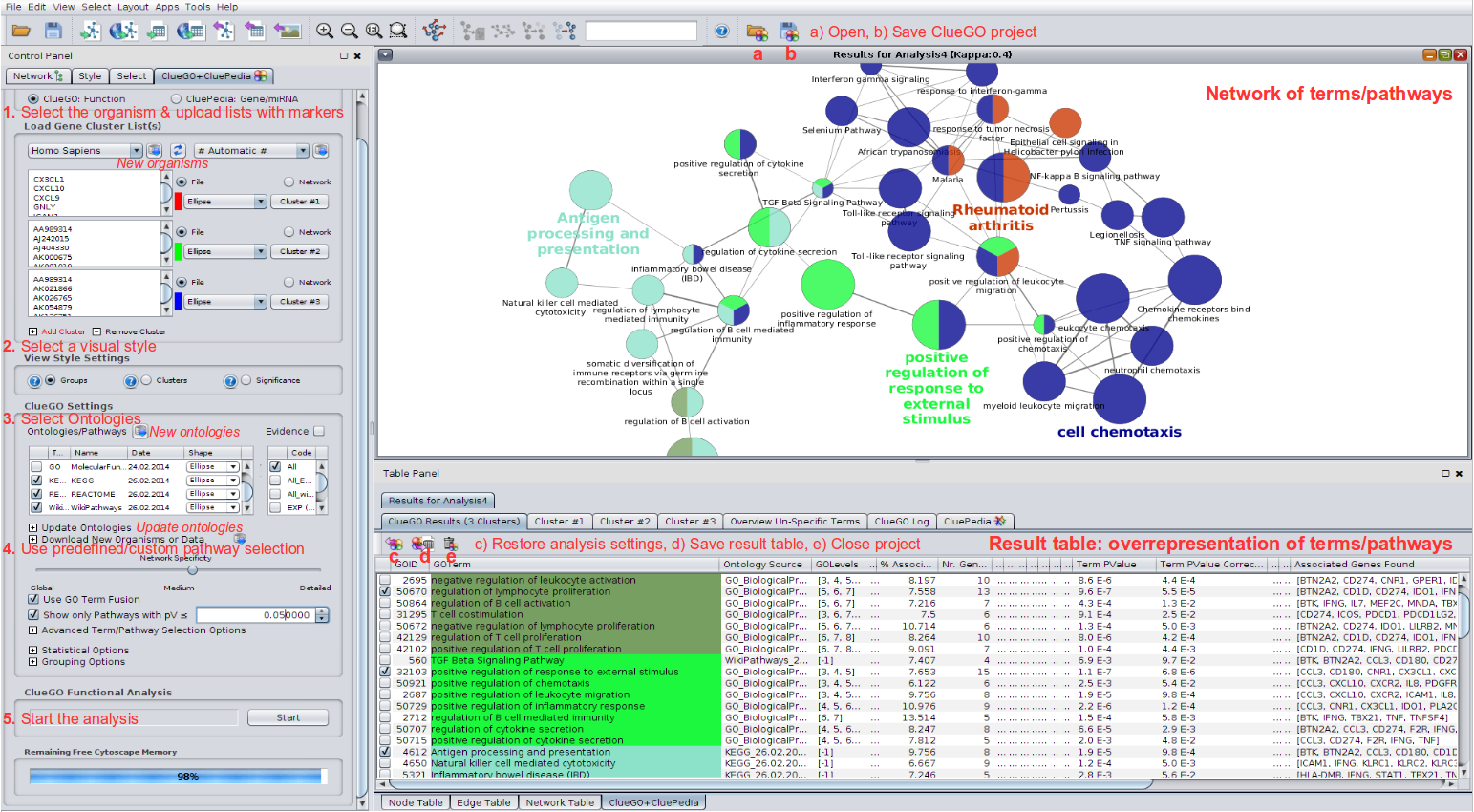
The merged network structure is normalized to guarantee a consistent and uniform description of the network nodes and edges and to enrich all integrated data with additional annotations retrieved from genome-wide databases like NCBI, thus producing a pathway fully manageable through the Cytoscape environment. Moreover, ReNE allows researchers to merge multiple pathways coming from different sources. ReNE can automatically import a network layout from the Reactome or KEGG repositories, or work with custom pathways described using a standard OWL/XML data format that the Cytoscape import procedure accepts. In this context, we introduce ReNE, a Cytoscape 3.x plugin designed to automatically enrich a standard gene-based regulatory network with more detailed transcriptional, post transcriptional, and translational data, resulting in an enhanced network that more precisely models the actual biological regulatory mechanisms. To integrate networks with post transcriptional regulatory data, researchers are therefore forced to manually resort to specific third party databases. Regulatory networks are commonly limited to gene entities. Despite post transcriptional regulatory elements (i.e., miRNAs) are widely investigated in current research, their usage and visualization in biological networks is very limited. One of the biggest challenges in the study of biological regulatory mechanisms is the integration, americanmodeling, and analysis of the complex interactions which take place in biological networks. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
